In southern China, the genetically improved slash pine (Pinus elliottii) plays a crucial role in timber and resin production, with new shoot density being a key growth trait. Current manual counting methods are inefficient and inaccurate. Emerging technologies such as UAV-based RGB imaging and deep learning (DL) offer promising solutions.
However, DL methods face challenges in global feature capture, necessitating additional mechanisms. Innovations like the Vision Transformer and its derivatives (e.g., TransCrowd, CCTrans) show potential in plant trait counting, offering simplified and more effective approaches for large-scale and accurate data processing. This technological evolution presents an opportunity for research in automated new shoot detection in slash pines, utilizing these advanced DL methodologies.
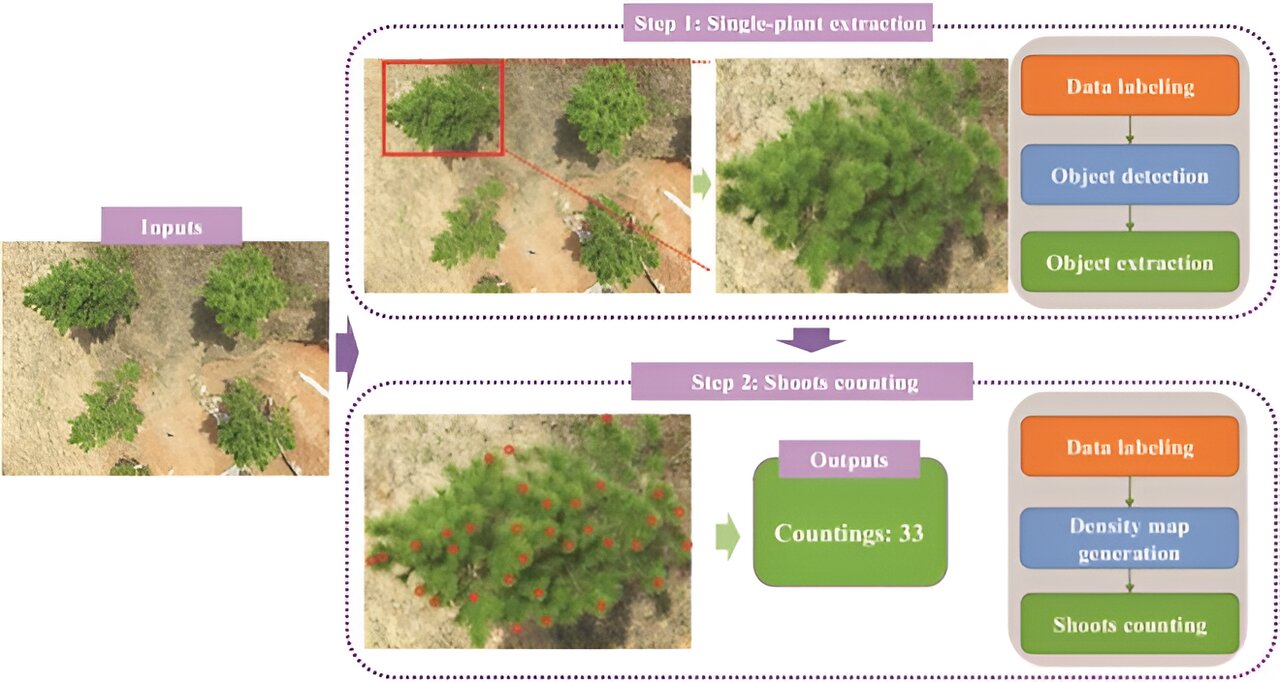
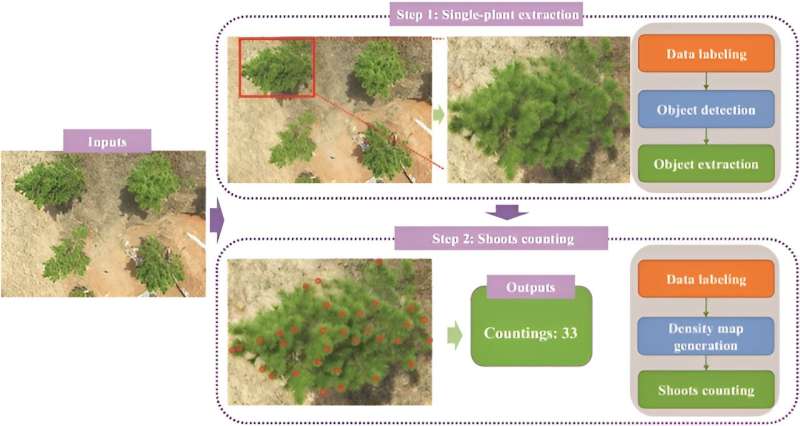
In July 2023, Plant Phenomics published a research article titled “CountShoots: Automatic Detection and Counting of Slash Pine New Shoots Using UAV Imagery.” This study introduces the Slash Pine Shoot Counting Network (SPSC-net), a model based on CCTrans, designed for counting new shoots of slash pine. It incorporates a feature pyramid module for accurate counting.
In the detection of slash pine trees, models like YOLOv5, Efficientnet, and YOLOX were compared, using a 0.5 threshold for tree identification. YOLOX demonstrated superior precision, recall, and average precision(AP), especially at a higher 0.75 threshold. In contrast, Faster-RCNN showed the lowest performance. Manual counting of 26 test images revealed that YOLOX had a lower false detection rate and EfficientNet had minimal missed targets.
YOLOX excelled in complex and overlapping target scenarios. For the detection of new shoots, the study compared balanced and unbalanced OT frameworks while assessing different transposition cost matrices. The perspective-guided model displayed the best performance, validating the efficacy of nonequilibrium OT for density regression. SPSC-net achieved the lowest MSE and MAE among all models, outperforming DM-Count, CSR-net, and MCNN. Scatter plots and density maps demonstrated the high prediction accuracy of the SPSC-net.
On this basis, the study developed CountShoots, a system of extracting and counting slash pine. Implemented on the Flask framework, it features modules for user interaction, model loading, plant extraction, and shoot counting. The process involves uploading images, extracting plant data, counting shoots, and providing feedback on the results, all streamlined for user convenience. The study confirmed the effectiveness of the SPSC-net in multiscale image processing of slash pine.
YOLOX and SPSC-net were compared with other models, demonstrating superior detection and counting accuracy. SPSC-net’s self-attention mechanism and feature pyramid fusion enable detailed and semantically rich feature extraction. Despite its success, there are limitations to consider, such as potential obstruction from the canopy layer and restrictions on UAV flight height.
In conclusion, the research developed a comprehensive pipeline using SPSC-net and YOLOX for accurate slash pine shoot counting and crown detection, offering a robust tool for forestry research and genetic breeding of slash pine.
More information:
Xia Hao et al, CountShoots: Automatic Detection and Counting of Slash Pine New Shoots Using UAV Imagery, Plant Phenomics (2023). DOI: 10.34133/plantphenomics.0065
Provided by
NanJing Agricultural University
Citation:
‘CountShoots’ unveils advanced UAV and AI techniques for precise slash pine shoot counting (2023, December 16)
retrieved 18 December 2023
from https://phys.org/news/2023-12-countshoots-unveils-advanced-uav-ai.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.





