Over the past decade, generative deep learning models have been applied successfully to the design of novel drug molecules, organic synthesis routes, and functional molecules tailored for electronic/optoelectronic devices. This is largely enabled by the availability of SMILES representation for molecules—an invertible and invariant representation well-suited for natural language processing models like recurrent neural networks, transformers, etc.
However, designing crystalline inorganic solids with desired properties remains a formidable challenge. This is primarily due to the lack of a “SMILES equivalent” crystal representation to bridge periodic solid-state materials and state-of-the-art deep learning architectures.
Previous methods for inverse crystal design mostly relied on 3D voxel grids or absolute spatial coordinates to represent structures. But these approaches intrinsically lack rotational invariance. There are also attempts at using crystal graphs, which are invariant but not invertible due to the absence of explicit periodicity or composition information. To address this challenge, we proposed a new crystal representation called SLICES. The study is published in the journal Nature Communications.
The core idea behind SLICES
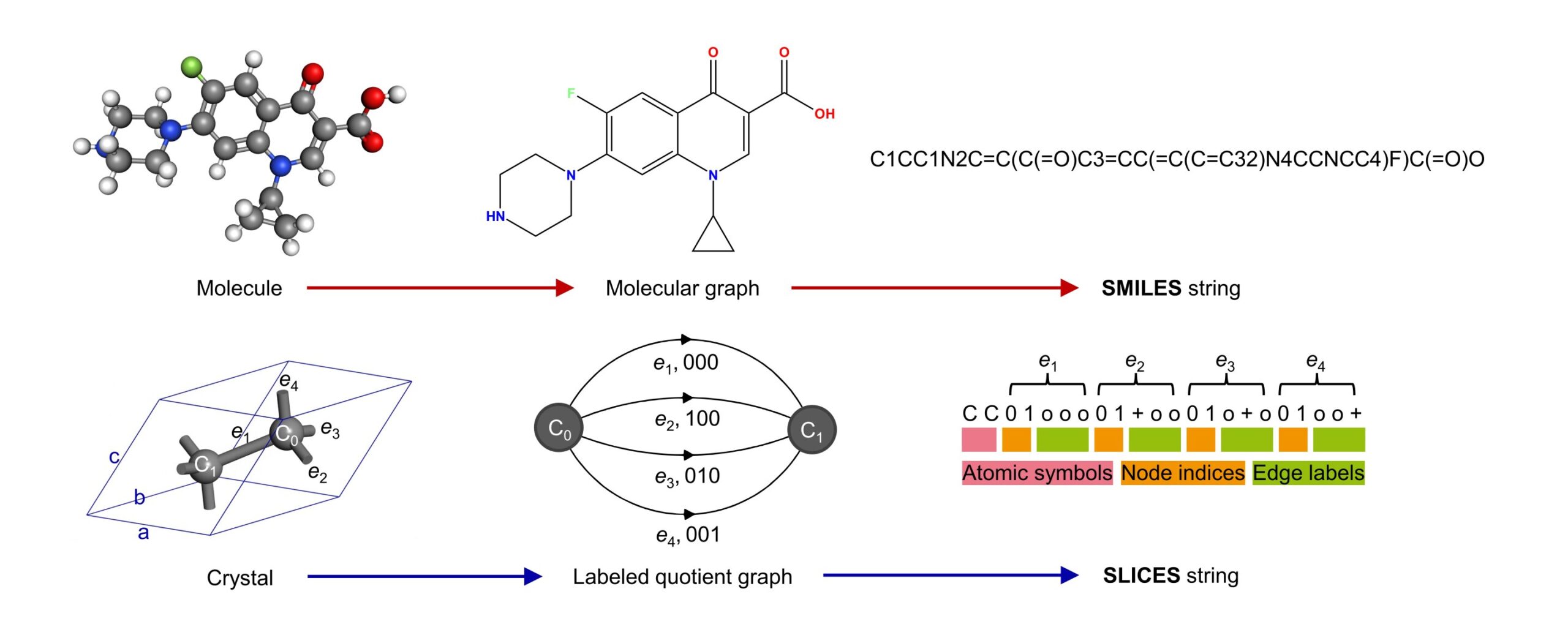
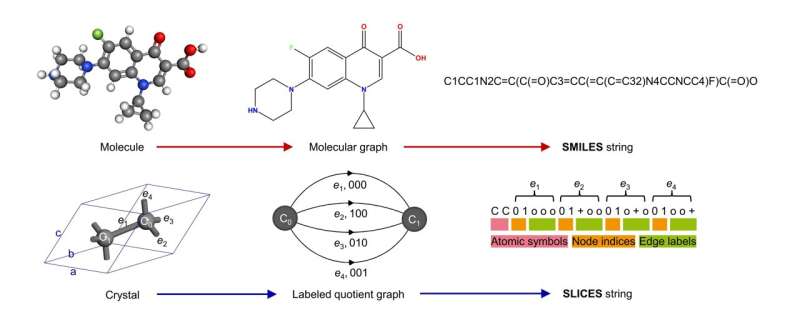
The key motivation behind developing SLICES is to create a crystal representation that is invertible and invariant, analogous to the SMILES representation used widely for molecular inverse design (Figure 1). Invertibility means the representation can be unambiguously converted back to the original crystal structure. This is essential for generative models to conduct inverse design, where the models create new crystal structures that are decoded from the representation.
Invariance indicates the representation remains unchanged under translations, rotations, and permutations of the crystal structure. Satisfying invariances allows the representation to purely focus on encoding the essential topological and compositional information of a system rather than superficial features that change under transformations. This reduces redundancy and improves learning efficiency.
By satisfying invertibility and invariances, SLICES enables efficient exploration of the vast chemical compound space for crystalline materials using deep generative models.
How SLICES represents crystals
Conceptually, SLICES encodes the topology and composition of crystal structures into strings, much like how SMILES converts molecular graphs into line notations. More specifically, SLICES leverages the mathematical concept of “labeled quotient graphs” to represent periodic crystal structures. The atoms and bonds within a unit cell are mapped to nodes and edges of the quotient graph. Additional labels are assigned to edges indicating the periodic shift vectors required to connect equivalent atoms in neighboring unit cells.
An example is the crystal structure of diamond (Figure 1), which contains two carbon atoms bonded together in the primitive unit cell. The SLICES string explicitly encodes the atomic symbols “C” and the edge label “001” denoting the periodic bond that propagates along the [001] direction. By parsing the SLICES string, both the composition and connectivity of the diamond structure can be obtained.
Notably, SLICES only encodes topology and composition information. Attributes like atomic coordinates and lattice parameters are not explicitly embedded. This makes SLICES invariant to translations, rotations, and atom index permutations by design.
Reconstructing crystal structures from SLICES
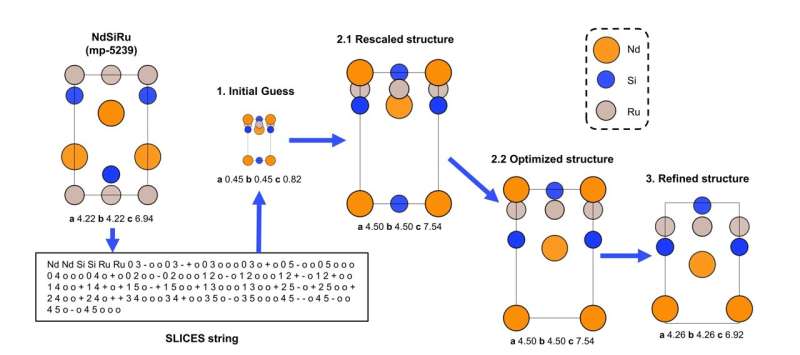
While encoding crystals into SLICES is relatively straightforward, the challenge lies in ensuring invertibility—the ability to accurately rebuild crystal structures from the SLICES strings. To achieve invertibility, we developed a reconstruction pipeline (Figure 2) for SLICES that contains three key steps:
- Generate an initial structure using graph theory techniques based on the topology and connectivity information parsed from the input SLICES string.
- Optimize the initial structure to have chemically reasonable geometry using a modified interatomic potential.
- Further refine the structure with a graph neural network-based universal crystal relaxation model.
The reconstruction performance was benchmarked on a database containing more than 40,000 experimentally known materials with up to 20 atoms per unit cell. The reconstruction pipeline for SLICES was able to reconstruct 94.95% of the original structures, substantially outperforming previous methods. This invertibility of SLICES allows for the generation of new structures from learned representations, which is key to inverse materials design.
Application in inverse design of functional materials
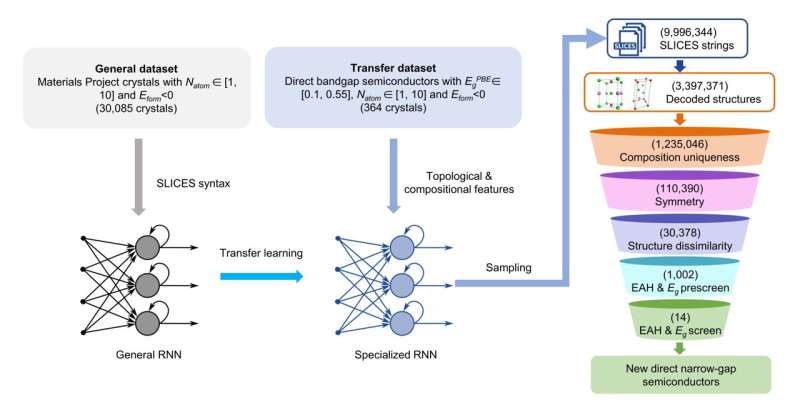
As a demonstration, we applied SLICES in the inverse design of direct narrow-bandgap semiconductors for optoelectronic devices using recurrent neural networks (RNN). The workflow consists of (Figure 3):
- Training an RNN model on known crystal structures to learn the underlying SLICES syntax and composition/topology features that correlate with targeted electronic properties.
- Using the trained RNN to generate hypothetical SLICES strings.
- Reconstructing the SLICES strings into crystal structures.
- Screening the structures using ab initio calculations and AI models to identify candidates that meet the design criteria.
Through this workflow combining SLICES, RNN, and high-throughput computations, 14 novel semiconductors with direct bandgaps in the optimal range were discovered (Figure 4). This showcases the promise of SLICES as an enabler for accelerated discovery of functional materials using generative AI.
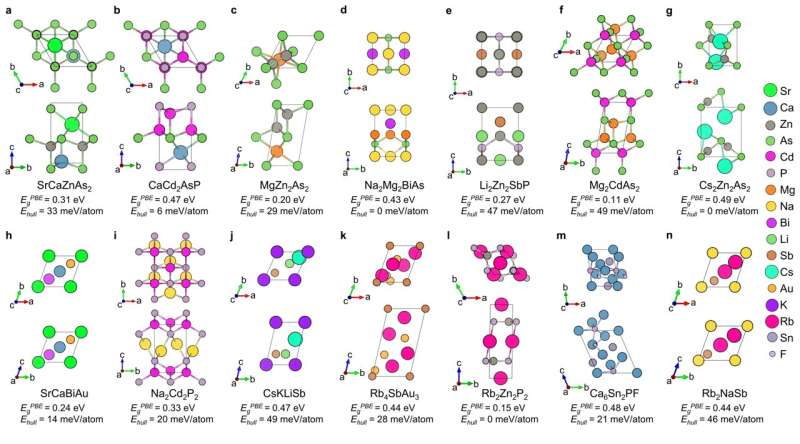
Directed generation of new materials with specified formation energies
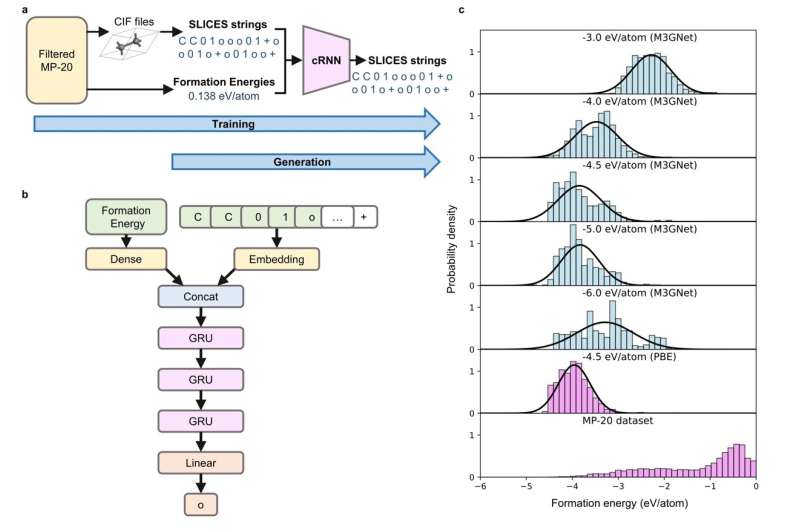
In addition, we employ a conditional recurrent neural network (cRNN) architecture, as illustrated in Figure 5, to generate SLICES strings corresponding to crystals with a desired formation energy specified by the user. The distribution of formation energies of the generated structures shifts closer to the specified target value relative to the dataset distribution. SLICES-based cRNN significantly outperform previous state-of-the-art models. This approach marks a significant advancement in the ability to design and discover new materials in a controlled and precise manner.
As the first string-based invertible and invariant crystal representation, SLICES opens up many exciting opportunities in the inverse design of crystalline solids, just as SMILES has done for molecules in the past decade. Just in the past few years, we have witnessed tremendous advances in generative models ranging from images, videos, speech, to proteins and molecules. We envision solid materials being the next frontier, thanks to this new capacity for data-efficient, chemistry-integrated exploration empowered by representations like SLICES.
This story is part of Science X Dialog, where researchers can report findings from their published research articles. Visit this page for information about ScienceX Dialog and how to participate.
More information:
Hang Xiao et al, An invertible, invariant crystal representation for inverse design of solid-state materials using generative deep learning, Nature Communications (2023). DOI: 10.1038/s41467-023-42870-7
Hang Xiao is affiliated with the School of Interdisciplinary Studies, Lingnan University; he earned his PhD from Columbia University. Yan Chen is affiliated with the Laboratory for Multiscale Mechanics and Medical Science, SV LAB, School of Aerospace, Xi’an Jiaotong University, where he also earned his PhD.
Citation:
Crystal language empowers AI to design novel materials with desired properties (2023, December 13)
retrieved 13 December 2023
from https://phys.org/news/2023-12-crystal-language-empowers-ai-materials.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.





