Protein-splitting enzymes play an important role in many physiological processes. Such proteases are generally present in an inactive state, only becoming activated under certain conditions. Some are linked to diseases like infections or cancer, making it important to have methods that can selectively detect active proteases.
In an article published in the journal Angewandte Chemie International Edition, scientists have introduced a new class of protease-activity sensors: gold nanoparticles equipped with peptide DNA.
Led by Devleena Samanta and Anna Capasso (The University of Texas at Austin, U.S.), the team has shown that these nanoprobes can sense multiple active proteases in parallel (multiplexed measurement). The method works at room temperature and does not require complicated sample preparation or elaborate instruments.
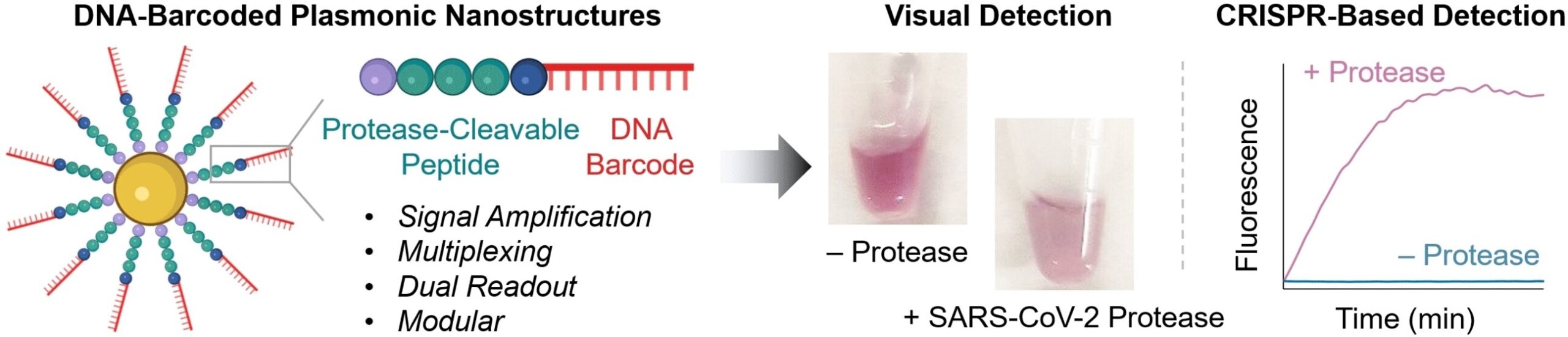
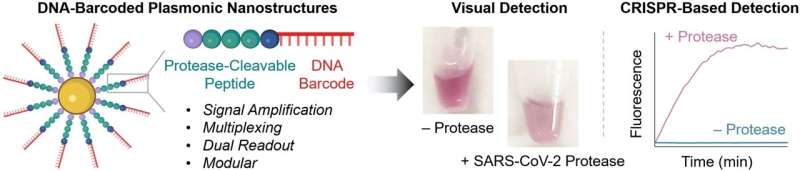
At the core of the novel probes are gold nanoparticles equipped with chains made of a peptide and a DNA fragment. The peptide structure is designed to be one that is split by the protease being detected. The DNA acts as a unique barcode for identifying the peptide and also amplifies the signal. If the desired protease is present in its active form in the sample, the peptide splits it. This releases the DNA barcode into the solution, where it can be detected based on its sequence.
To carry out this detection, the team uses a CRISPR/Cas12a test: the enzyme Cas12a is bound to a guide RNA (gRNA) to form an inactive complex. The gRNA contains a segment that specifically binds to the barcode DNA. This activates the Cas12a, so that it can now “cut up” single-stranded DNA (ssDNA).
For the test, the researchers add ssDNA molecules with a fluorescing group (fluorophore) at one end and a quencher, which “switches off” the fluorescence of the fluorophore (as long as they are close enough), at the other. If the ssDNA is cut up, the fluorophore and quencher move further apart. This results in strong fluorescence that indicates that the protease being tested for is present (detection limit of about 58 pM).
If no instruments are available on site and the test must go fast, detection is possible with the naked eye: if the protease splits the peptide on the probe, the surface charge of the gold nanoparticles changes and they aggregate. The color of these so-called “plasmonic nanostructures” depends significantly on their degree of aggregation. It is possible to detect nanomolar protease concentrations based on the color change in the test solution.
Multiplexed detection of the proteases 3CL and caspase3 allowed the team to demonstrate the high sensitivity and selectivity of their new method. 3CL is a marker for active coronavirus infection and COVID patients often also have elevated activity of the apoptosis marker caspase3. The clinical potential of this test was also demonstrated by the detection of cathepsin B, a protease related to colorectal cancer, in three different tumor cell lines obtained from patients.
These nanoprobes yield 100-fold higher fluorescence signals compared to commercial fluorescence-based protease sensors. Moreover, virtually any protease can be detected if the peptide it splits is known. Taken together, these nanoprobes can potentially enable early disease detection and improve the precision and reliability of diagnostic tests through multiplexing.
More information:
Subrata Pandit et al, DNA‐Barcoded Plasmonic Nanostructures for Activity‐Based Protease Sensing, Angewandte Chemie International Edition (2023). DOI: 10.1002/anie.202310964
Citation:
A nanoprobe with a barcode: Sensors detect active proteases (2023, December 12)
retrieved 12 December 2023
from https://phys.org/news/2023-12-nanoprobe-barcode-sensors-proteases.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.





