Tomato is a widely cultivated crop, valued for both culinary and medicinal purposes. Its vulnerability to various pests and diseases, especially affecting leaves, poses a challenge for growers. Traditional methods of disease identification, based on subjective human judgment, have proved inefficient and unreliable.
The advent of image processing technology, particularly deep learning, has revolutionized disease detection in agriculture. These techniques involve collecting and processing disease images, extracting features, and training models for accurate identification; despite advancements, challenges persist, such as accurately detecting small or blurred disease symptoms.
Researchers have developed several methods to overcome these limitations, including optimizing models and employing advanced algorithms. However, deep learning in plant disease recognition still encounters challenges like complexity and adaptability to diverse agricultural settings, directing ongoing research toward enhancing these technologies.
In May 2023,Plant Phenomics published a research article titled “An Effective Image-Based Tomato Leaf Disease Segmentation Method Using MC-UNet“.
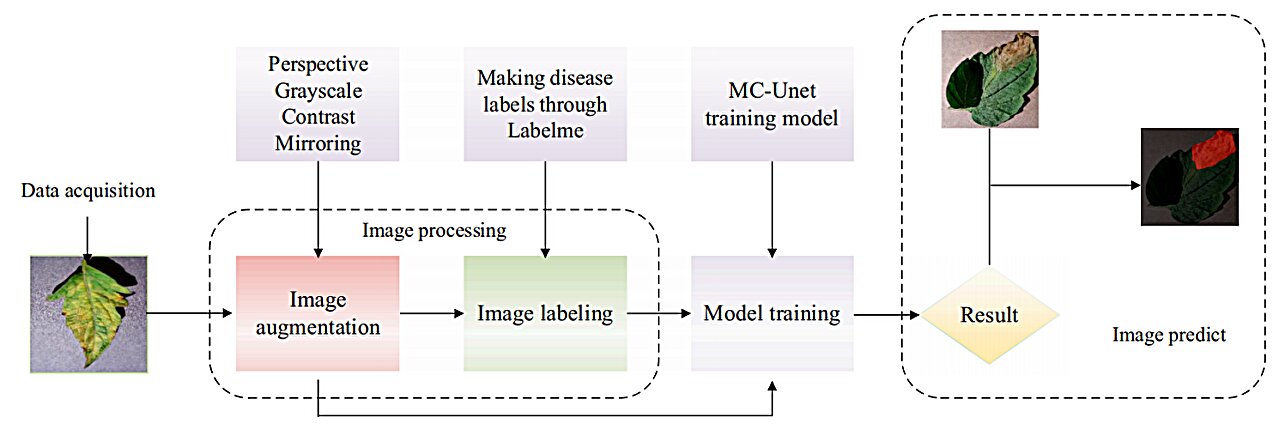
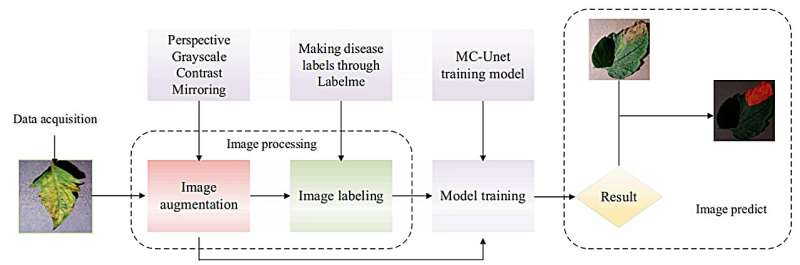
The research introduces the Cross-layer Attention Fusion Mechanism combined with a Multi-scale Convolution Module (MC-UNet), an enhanced image-based tomato leaf disease segmentation method based on UNet. This method incorporates a Multi-scale Convolution Module for obtaining multiscale information about tomato disease utilizing convolution kernels of various sizes and emphasizing edge features.
The Cross-layer Attention Fusion Mechanism is employed to pinpoint disease locations on tomato leaves, using SoftPool for information retention and the SeLU function to prevent neuron dropout. MC-UNet demonstrated 91.32% accuracy and 6.67M parameters in a self-built dataset, affirming its effectiveness for tomato leaf disease segmentation.
The research showed that MC-UNet outperforms other models in tomato disease segmentation. The effectiveness of individual modules like the Multi-scale Convolution Module (MCM) and Cross-layer Attention Fusion Mechanism (CAFM) was analyzed. MCM was efficient in extracting feature information, and CAFM improved the model’s performance by enhancing multiscale output fusion.
The use of SoftPool as a pooling layer exhibited a notable improvement in network performance compared to the original MaxPool in UNet. Ablation experiments confirmed the effectiveness of each module, with CAFM achieving the most significant improvement. MC-UNet’s robustness in different lighting environments was also evaluated, showing its superior performance over the baseline network UNet, especially in tiny and edge-blurred leaf diseases.
The experiment concluded that MC-UNet is a suitable model for tomato leaf disease segmentation. It significantly outperforms other networks in accuracy and has strong generalization ability. However, it showed limitations in dealing with complex backgrounds, indicating the necessity for future research focused on multistage segmentation models and datasets with complex backgrounds to enhance the model’s resistance to interference.
More information:
Yubao Deng et al, An Effective Image-Based Tomato Leaf Disease Segmentation Method Using MC-UNet, Plant Phenomics (2023). DOI: 10.34133/plantphenomics.0049
Provided by
NanJing Agricultural University
Citation:
Transforming tomato crop health: Introducing a method for advanced leaf disease detection and segmentation (2023, December 11)
retrieved 11 December 2023
from https://phys.org/news/2023-12-tomato-crop-health-method-advanced.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.





