
The building blocks for new drugs that help fight bacteria that are resistant to known antibiotics, for example, should be as cost-effective and environmentally friendly as possible. Enzymes are ideal for this purpose. For example, they can produce or combine different components of active substances.
In his master’s thesis in the Microbial Biotechnology group at Ruhr University Bochum, Simon Schröder characterized an enzyme in more detail that is capable of forming a desired nitrogen–nitrogen bond in molecules. He also found other enzymes that can do this. The work is published in the journal Molecular Catalysis.
Building blocks limit the design of new active substances
Researchers are in constant competition with harmful microorganisms that develop antibiotic resistance. In the search for new active substances, they traditionally try to isolate microorganisms from nature that exhibit antibiotic behavior. They then identify the substances responsible and study their function. Today, this process is complemented by computer-aided methods that make it possible to design tailor-made new molecules that have specific effects on organisms and their metabolic processes.
“However, the design and production of such artificial compounds is often limited by which precursor molecules or building blocks are available for their production,” explains Schröder. Ideally, their production process should be economical and ecological, for example by using microorganisms or their catalytic enzymes. The expansion of the modular system of available molecules to produce new drugs is therefore correspondingly important and interesting.

Making the desired bond more easily accessible
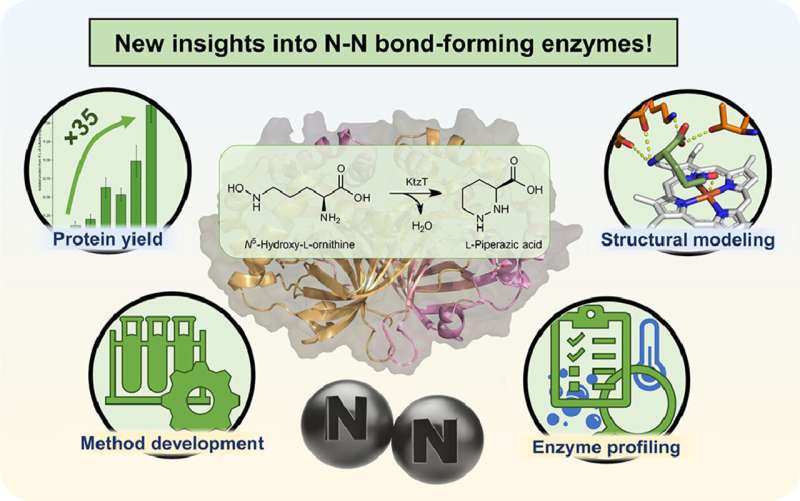
“We are working on the production of a specific type of such molecules,” explains Schröder. In 2017, an enzyme was isolated that can form the nitrogen-nitrogen bond in molecules, which is rarely found in nature. However, very little is still known about this enzyme with the systematic name “KtzT”: How does it work? In which compounds can it form this bond? Is it suitable to produce pharmaceutically relevant molecules?
“Initially, we were able to improve the production and isolation of this enzyme in the laboratory by a factor of 35,” reports Schröder. “This enabled us to characterize KtzT, i.e., to identify its optimal reaction conditions: At what temperature, what pH value does it work best and how stable is it under a wide range of conditions?”
The research team has also found and isolated KtzT-like enzymes and shown that they are also able to catalyze the reaction. “We were also able to implement a multi-step reaction with several enzymes, making the nitrogen-nitrogen bond even easier to access,” says Schröder.
Among other things, he used bioinformatic methods to develop a structural model of the enzyme, which enables hypotheses to be made regarding the reaction mechanism and the enzyme to be specifically modified so that it can also form the nitrogen-nitrogen bond in other compounds.
More information:
Simon Schröder et al, Enhancing biocatalytical N–N bond formation with the actinobacterial piperazate synthase KtzT, Molecular Catalysis (2023). DOI: 10.1016/j.mcat.2023.113733
Provided by
Ruhr-Universitaet-Bochum
Citation:
Researcher characterizes enzymes with N–N bonds for antibacterial applications (2023, December 8)
retrieved 8 December 2023
from https://phys.org/news/2023-12-characterizes-enzymes-nn-bonds-antibacterial.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.





